|
In our laboratory, acoustic shearing with a Covaris instrument (Covaris, Woburn, MA) is typically done to obtain DNA fragments in the 100–5000 bp range, while Covaris g-TUBEs are employed for the 6–20 Kbp range necessary for mate-pair libraries. DNA fragmentation is typically done by physical methods (i.e., acoustic shearing and sonication) or enzymatic methods (i.e., non-specific endonuclease cocktails and transposase tagmentation reactions)( 12). Three approaches are available to fragment nucleic acid chains: physical, enzymatic, and chemical. The size of the target DNA fragments in the final library is a key parameter for NGS library construction. In general, the core steps in preparing RNA or DNA for NGS analysis are: ( i) fragmenting and/or sizing the target sequences to a desired length, ( ii) converting target to double-stranded DNA, ( iii) attaching oligonucleotide adapters to the ends of target fragments, and ( iv) quantitating the final library product for sequencing. Cell photograph courtesy of Annie Cavanagh, Wellcome Images. One difference between the two technologies is that Illumina allows sequencing from both ends of the library insert (i.e., paired end sequencing). The final step generates the actual sequence via the chemistries for each technology. The next step involves clonal amplification of the library, by either cluster generation for Illumina or microemulsion PCR for Ion Torrent. DNA Fragments are converted into the library by ligation to sequencing adapters containing specific sequences designed to interact with the NGS platform, either the surface of the flow-cell (Illumina) or beads (Ion Torrent). RNA is converted to cDNA by reverse transcription. RNA or DNA is extracted from sample tissue/cells and fragmented. Basic workflow for NGS library preparation. However, it should be noted that almost all of the principles discussed in this review can be applied with minimal modification to NGS platforms developed by Life Technologies, Roche, and Pacific Biosciences. 
Here, we compare and contrast various library preparation strategies and NGS applications, focusing primarily on those compatible with Illumina sequencing technology. 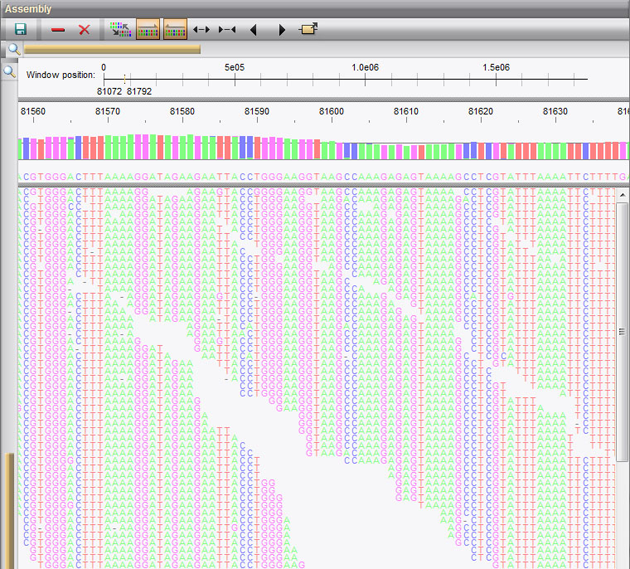
For example, NGS library preparation has now been successfully demonstrated for sequencing RNA and DNA from single cells ( 3–11).įundamental to NGS library construction is the preparation of the nucleic acid target, RNA or DNA, into a form that is compatible with the sequencing system to be used ( Figure 1). During this time, as sequencing technologies have improved and evolved, so too have methods for preparing nucleic acids for sequencing and constructing NGS libraries ( 1, 2). 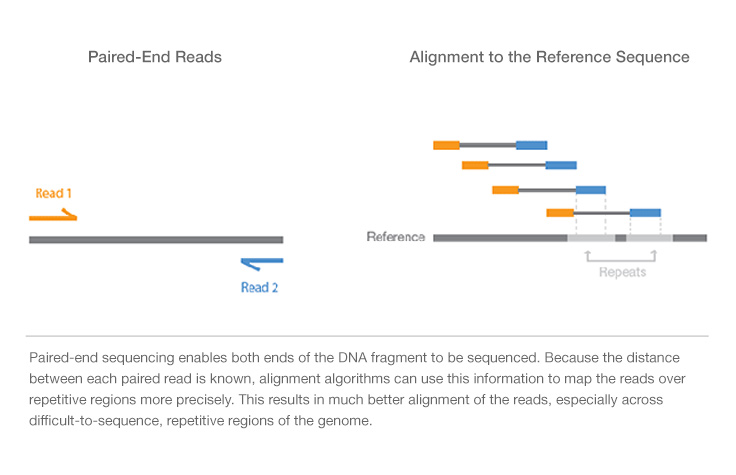
Over the past five years, next-generation sequencing (NGS) technology has become widely available to life scientists.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
